
In May 2024, the biotechnology world was once again transformed with the release of AlphaFold 3, developed by Google DeepMind and Isomorphic Labs. This new artificial intelligence (AI) model promises to revolutionize biomolecular research and drug development, significantly expanding the capabilities of its previous versions.1,2
What Is AlphaFold 3?
AlphaFold 3 is an AI model designed to predict the three-dimensional structure of proteins and other biomolecules, including DNA, RNA, and small molecules such as ligand-proteins—substances that bind specifically to a receptor or another molecule, usually to trigger a biological response.1,2
In addition to predicting the shapes of proteins, AlphaFold 3 can create models of complex interactions between different types of molecules, offering a more detailed understanding of biological processes.1,2
Why Is Modeling Complex Interactions Between Molecules Important for Discovering New Treatments?
Proteins and other molecules play essential roles in the body—they are fundamental to the structure, function, and regulation of tissues and organs. These molecules have complex three-dimensional forms that, in the case of proteins, are determined by their sequence of amino acids. How these sequences “fold” or are structured is crucial for their function.3,4
Each protein folds in a specific way, forming a unique three-dimensional structure. This folding process is influenced by intramolecular interactions and chemical forces such as:4
- hydrogen bonds,
- hydrophobic interactions,
- positive and negative electric charges in the side chains of amino acids.
The electric charges on proteins can attract or repel other molecules, determining how and where those proteins will bind to other molecules.4
For example, hemoglobin is an essential protein for transporting oxygen in the blood. Its three-dimensional structure allows it to bind oxygen in the lungs and release it in body tissues. Any alteration in the shape of hemoglobin—such as mutations that cause sickle-cell anemia—can impair its ability to transport oxygen efficiently.5

Figure 1. Illustration of the red blood cell and the three-dimensional hemoglobin molecule. Own image. All rights reserved.
Understanding the three-dimensional shape of a molecule is different from knowing only its amino acid sequence, because a molecule’s function depends on its specific shape. The amino acid sequence gives the code, but it is the resulting three-dimensional form that defines how the protein interacts with other molecules.4
This understanding is fundamental for discovering new treatments, allowing scientists to design drugs that can bind specifically to target proteins, modulating their functions to treat diseases.4
Moreover, the three-dimensional conformation of proteins is critical to understanding their functional interactions and internal dynamics— aspects difficult to capture with just the linear amino acid sequence. Thus, advanced methods like AlphaFold help reveal intricate details of protein-protein interactions, essential to biological processes such as cell signaling and metabolism.6
Advances in predicting and designing protein structures
Predicting the three-dimensional structure of proteins from amino acid sequences has been a major challenge in computational biophysics for decades. More recently, the inverse problem—that is, designing a sequence of amino acids that will fold into a specific three-dimensional structure—has been receiving growing attention as a potential path toward rational engineering of proteins with useful functions in biotechnology and medicine.4
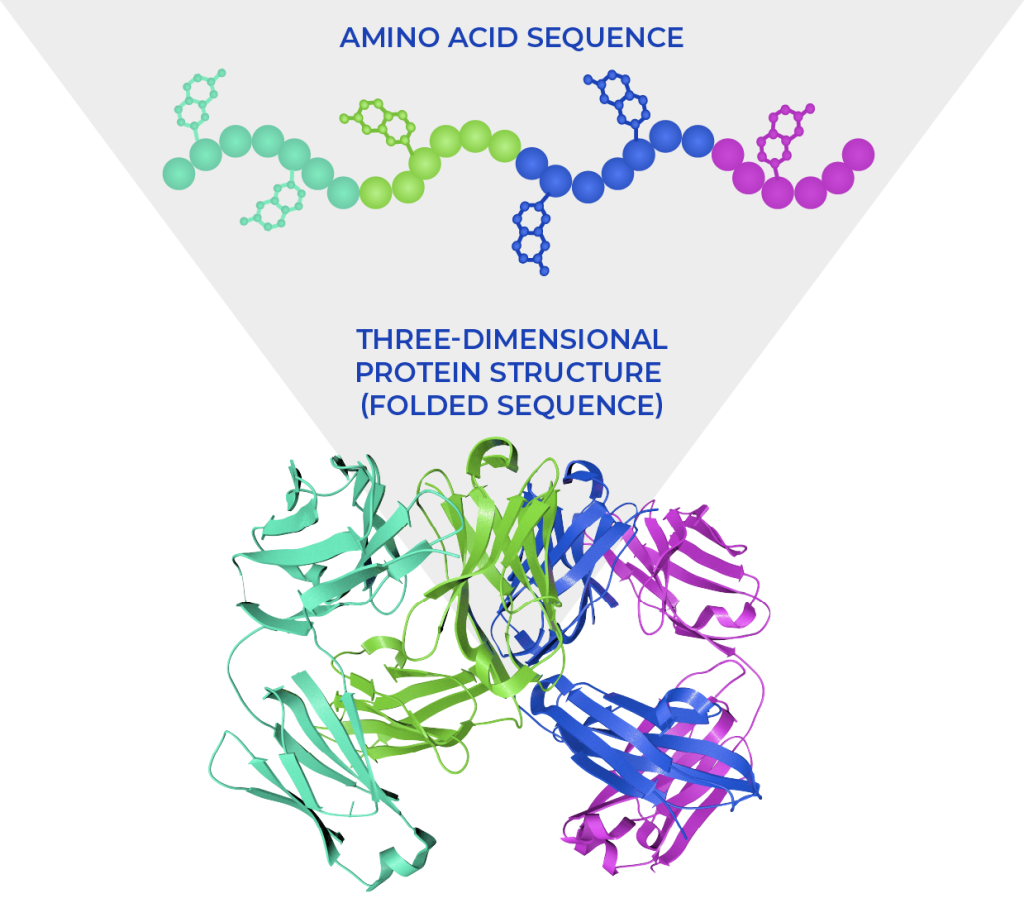
Figure 2. Representation of the linear amino acid sequence and the three-dimensional structure assumed by the protein, with specific “folds.” Own image. All rights reserved.
The sharp advances made in the last decade in predicting and designing protein structures have been driven mainly by:
- increases in computational power,
- the rapid growth of protein sequence and structure databases.
These have enabled the creation of new algorithms to design protein folds and protein-protein interfaces, which have been used to design novel high-complexity models with new or enhanced properties, as well as signaling proteins with therapeutic potential.4
Importance for Pharmaceutical Research & Development
The ability of AlphaFold 3 to accurately predict structures and molecular interactions has profound implications for drug discovery. Here are some of the main benefits:1,2
- Identification of Therapeutic Targets: With AlphaFold 3, researchers can identify new molecular targets for drug development more quickly and accurately, which can accelerate the discovery of treatments for complex diseases.
- More Effective Drug Design: Accurate prediction of interactions between proteins and other molecules allows for the creation of drugs that bind more effectively to their targets, increasing efficacy and reducing side effects.
- Acceleration of Clinical Development: By providing detailed structural models, AlphaFold 3 facilitates the preclinical development phase, allowing researchers to move more quickly into clinical trials.
Representation of a molecule binding to its specific cellular receptors and being internalized into the cell. Own source. All rights reserved.
Benefits of AlphaFold 3
- Improved Accuracy: AlphaFold 3 shows at least a 50% improvement in predicting molecular interactions compared to existing methods, and in some cases, it has doubled accuracy.1,2
- Free Access for Research: Although the commercial model is managed by Isomorphic Labs, AlphaFold 3 offers free access for non-commercial uses, democratizing biomolecular research.1,2
- Collaboration with the Pharmaceutical Industry: Isomorphic Labs is already collaborating with pharmaceutical companies to apply AlphaFold 3 to real drug design challenges, aiming to develop new treatments that could transform patients’ lives.1,2
Why This Matters
The computational advances in developing systems and algorithms that can perform complex tasks—including machine learning and analysis of large volumes of data—combined with the growth in available information, such as protein databases, represent not only a technological advance, but also a powerful tool that can transform the pharmaceutical industry’s approach to drug discovery. The ability to model complex interactions among biomolecules opens up new possibilities to understand and treat a variety of diseases, from cancer to infectious and genetic disorders.1,2,4
For pharmaceutical marketing and medical professionals, understanding the potential of AI models like AlphaFold 3 is extremely valuable, as this technology not only accelerates the development of new products, but also provides a significant competitive advantage in the global pharmaceutical market.2
In summary, AlphaFold 3 seems to be more than a technical update; it may represent a quantum leap in our ability to understand and manipulate molecular biology, with a direct and significant impact on global health.1,2
Let us follow how this unfolds in the years to come!
How to cite this article:
KACHI. A New Revolution in Drug Discovery? Impact on the Pharmaceutical Industry. São Paulo: KACHI Comunicação Científica, Dia Mês Ano. Disponível em: https://kachi.com.br/en/trends-in-health-communication-insights-for-modern-pharma-marketing.
References:
- Abramson J, Adler J, Dunger J, et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 2024;630(8016):493–500.
- AlphaFold 3 predicts the structure and interactions of all of life’s molecules [Internet]. Google. 2024 [cited 2024 Jun 15];Available from:https://blog.google/technology/ai/google-deepmind-isomorphic-alphafold-3-ai-model/
- Alberts B, Johnson A, Lewis J, Raff M, Roberts K, Walter P. Molecular Biology of the Cell. Garland Science; 2002.
- Kuhlman B, Bradley P. Advances in protein structure prediction and design. Nat Rev Mol Cell Biol 2019;20(11):681–97.
- Ahmed MH, Ghatge MS, Safo MK. Hemoglobin: Structure, Function and Allostery. Subcell Biochem 2020;94:345–82.
- Kryshtafovych A, Schwede T, Topf M, Fidelis K, Moult J. Critical assessment of methods of protein structure prediction (CASP)-Round XIII. Proteins 2019;87(12):1011–20.



